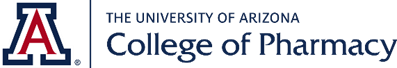Project 1: Drug Discovery and Pharmacology of SARS-CoV-2
To combat the COVID-19 pandemic, we joined the force with many other to develop antivirals against SARS-CoV-2. We are pursuing several drug targets including the main protease (Mpro or 3CLpro), the papain-like protease (PLpro), the nucleoprotein, and the polymerase. The goal is to identify broad-spectrum antiviral drug candidates with a high genetic barrier to drug resistance. For this, we relay on high-throughput screening and structure-based drug design to inform the lead optimization through iterative cycles of design, synthesis, and pharmacological characterization. We are also interested in profiling the genetic barrier to drug resistance of our leads and apply the knowledge to guide the design of resistance avoiding antivirals. We recently made progress in discovering several promising Mpro inhibitors including boceprevir, GC-376, calpain inhibitors II and XII. Boceprevir, calpain inhibiotrs II and XII represent novel Mpro inhibitors that are distinct from previously reported Mpro inhibitors such as GC-376 analogs. All four compounds showed potent enzymatic inhibition and cellular antiviral activity against Mpro and infectious SARS-CoV-2. In collaboration with Dr. Yu Chen's group at the University of South Florida we solved multiple X-ray crystal structures of Mpro with the hits we identified. These high-resolution X-ray crystal structures provides a foundation for the designing of next-generation of Mpro inhibitors.
To combat the COVID-19 pandemic, we joined the force with many other to develop antivirals against SARS-CoV-2. We are pursuing several drug targets including the main protease (Mpro or 3CLpro), the papain-like protease (PLpro), the nucleoprotein, and the polymerase. The goal is to identify broad-spectrum antiviral drug candidates with a high genetic barrier to drug resistance. For this, we relay on high-throughput screening and structure-based drug design to inform the lead optimization through iterative cycles of design, synthesis, and pharmacological characterization. We are also interested in profiling the genetic barrier to drug resistance of our leads and apply the knowledge to guide the design of resistance avoiding antivirals. We recently made progress in discovering several promising Mpro inhibitors including boceprevir, GC-376, calpain inhibitors II and XII. Boceprevir, calpain inhibiotrs II and XII represent novel Mpro inhibitors that are distinct from previously reported Mpro inhibitors such as GC-376 analogs. All four compounds showed potent enzymatic inhibition and cellular antiviral activity against Mpro and infectious SARS-CoV-2. In collaboration with Dr. Yu Chen's group at the University of South Florida we solved multiple X-ray crystal structures of Mpro with the hits we identified. These high-resolution X-ray crystal structures provides a foundation for the designing of next-generation of Mpro inhibitors.
Project 2: Drug Discovery and Pharmacology of Influenza Antivirals
The efficacy of current anti-influenza drugs is limited by the emergence of multidrug-resistant influenza viruses. Adamantanes (amantadine and rimantadine) are no longer recommended for prophylaxis and treatment of influenza infection. Resistance against neuraminidase inhibitors, oseltamivir, zanamivir, and paramivir, are continuously on the rise. As influenza viruses continuously undergo antigenic shift and antigenic drift, it is imperative to develop next generation of influenza antivirals with a novel mechanism of action. My lab is interested in developing broad-spectrum anti-influenza drugs with a high genetic barrier to drug resistance. We are exploring both viral proteins and host factors as drug targets. Our initial efforts have yielded several promising drug candidates including viral M2 channel blockers, viral polymerase PA-PB1 inhibitors, host kinase inhibitors, receptor antagonists, and protease inhibitors. Representative host-targeting antivirals we have discovered include cyclosporine A, dapivirine, hesperadine, and NMS-873.
With these novel lead compounds in hand, we are interested in understanding their mechanism of action and the mechanism of resistance. The approaches we have undertaken include in vitro resistance selection through serial viral passage, reverse genetics, and molecular modeling. We believe understanding the mechanism of drug resistance will help pave the way for the design of more effective and longer lasting antiviral drugs.
The efficacy of current anti-influenza drugs is limited by the emergence of multidrug-resistant influenza viruses. Adamantanes (amantadine and rimantadine) are no longer recommended for prophylaxis and treatment of influenza infection. Resistance against neuraminidase inhibitors, oseltamivir, zanamivir, and paramivir, are continuously on the rise. As influenza viruses continuously undergo antigenic shift and antigenic drift, it is imperative to develop next generation of influenza antivirals with a novel mechanism of action. My lab is interested in developing broad-spectrum anti-influenza drugs with a high genetic barrier to drug resistance. We are exploring both viral proteins and host factors as drug targets. Our initial efforts have yielded several promising drug candidates including viral M2 channel blockers, viral polymerase PA-PB1 inhibitors, host kinase inhibitors, receptor antagonists, and protease inhibitors. Representative host-targeting antivirals we have discovered include cyclosporine A, dapivirine, hesperadine, and NMS-873.
With these novel lead compounds in hand, we are interested in understanding their mechanism of action and the mechanism of resistance. The approaches we have undertaken include in vitro resistance selection through serial viral passage, reverse genetics, and molecular modeling. We believe understanding the mechanism of drug resistance will help pave the way for the design of more effective and longer lasting antiviral drugs.
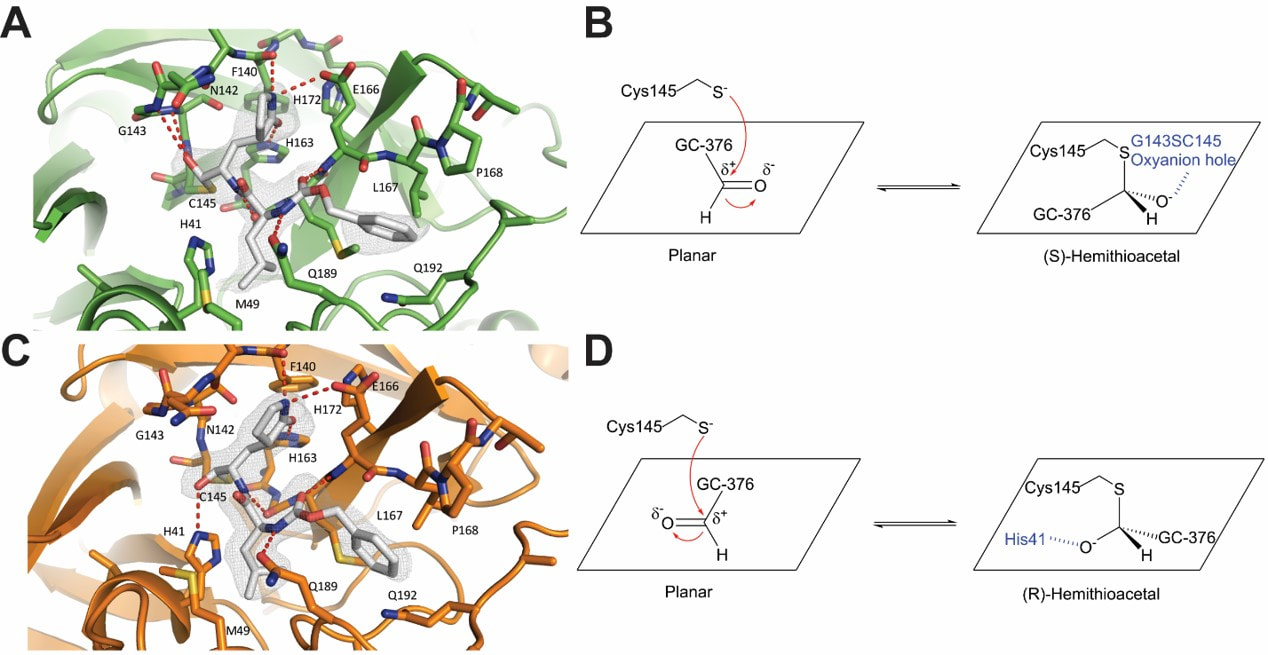
Rationally designed M2-S31N channel blockers with potent antiviral activity
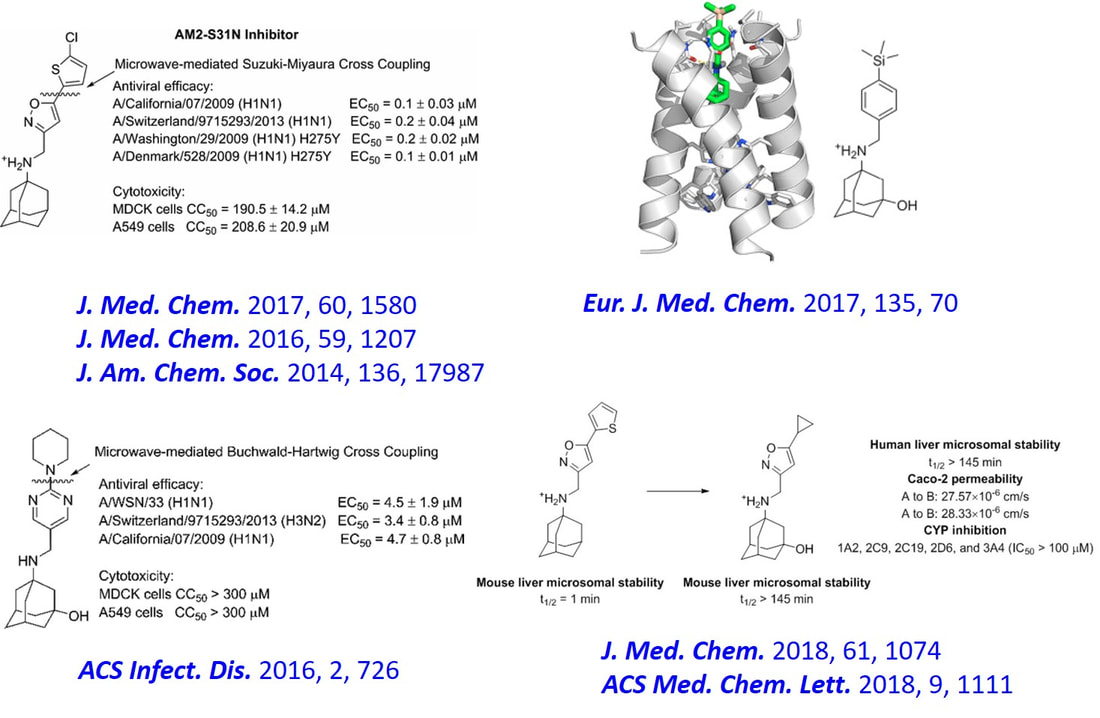
Rational designed tetrazole inhibitors targeting the influenza polymerase PA-PB1 protein protein interactions
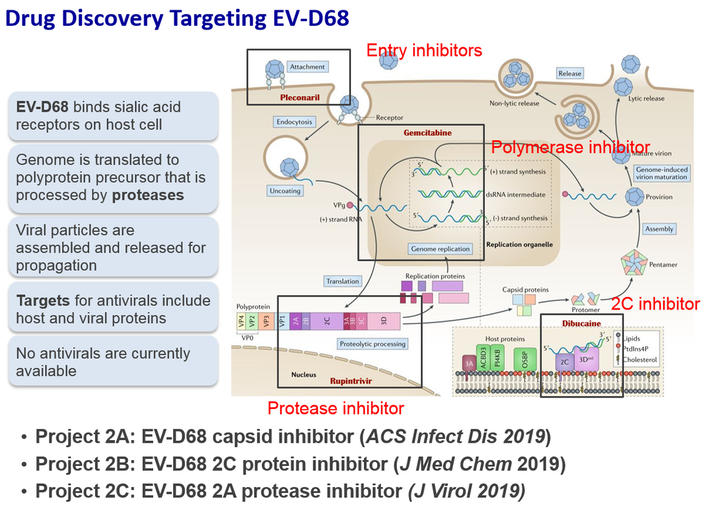
Project 3: Drug Discovery and Pharmacology of Enterovirus Antivirals
Enteroviruses are important human pathogens with disease manifestations ranging from mild respiratory illness to more severe neurological complications such as encephalitis, aseptic meningitis, acute flaccid paralysis/myelitis, and even death. The enterovirus genera of the Picornaviridae family contains poliovirus, rhinovirus, enterovirus A71 (EV-A71), enterovirus D68 (EV-D68), and coxsackievirus. Although poliovirus was nearly eradicated by vaccination, there are currently no antivirals or vaccines for the remaining non-polio enteroviruses. We are particularly interested in EV-D68, which has been recently labeled as "new ploliovirus" due to its link with neurological complications such as acute flaccid myelitis (AFM). Contemporary EV-D68 viruses appear to become more virulent and there is a biannual pattern of EV-D68 outbreak the U.S. As there is currently no antivirals available, we aim to develop antivirals against EV-D68 by targeting essential viral proteins such as the viral capsid protein VP1, the 2A protease, the 3C protease, the 2C protein, and the viral 3D polymerase. As EV-D68 is a RNA virus with an high mutation rate, we intend to develop a combination therapy approach to decrease the pace of drug resistance. For this, we will first demonstrate the in vivo antiviral efficacy of each of our antiviral drug candidate in the EV-D68 infection mouse model. Other than designing EV-D68 specific antivirals, we also made progress in developing broad-spectrum antivirals that target not only EV-D68, but also EV-A71, poliovirus, and coxsackievirus.
Enteroviruses are important human pathogens with disease manifestations ranging from mild respiratory illness to more severe neurological complications such as encephalitis, aseptic meningitis, acute flaccid paralysis/myelitis, and even death. The enterovirus genera of the Picornaviridae family contains poliovirus, rhinovirus, enterovirus A71 (EV-A71), enterovirus D68 (EV-D68), and coxsackievirus. Although poliovirus was nearly eradicated by vaccination, there are currently no antivirals or vaccines for the remaining non-polio enteroviruses. We are particularly interested in EV-D68, which has been recently labeled as "new ploliovirus" due to its link with neurological complications such as acute flaccid myelitis (AFM). Contemporary EV-D68 viruses appear to become more virulent and there is a biannual pattern of EV-D68 outbreak the U.S. As there is currently no antivirals available, we aim to develop antivirals against EV-D68 by targeting essential viral proteins such as the viral capsid protein VP1, the 2A protease, the 3C protease, the 2C protein, and the viral 3D polymerase. As EV-D68 is a RNA virus with an high mutation rate, we intend to develop a combination therapy approach to decrease the pace of drug resistance. For this, we will first demonstrate the in vivo antiviral efficacy of each of our antiviral drug candidate in the EV-D68 infection mouse model. Other than designing EV-D68 specific antivirals, we also made progress in developing broad-spectrum antivirals that target not only EV-D68, but also EV-A71, poliovirus, and coxsackievirus.
Project 4: Drug Resistance Mechanism of Antivirals
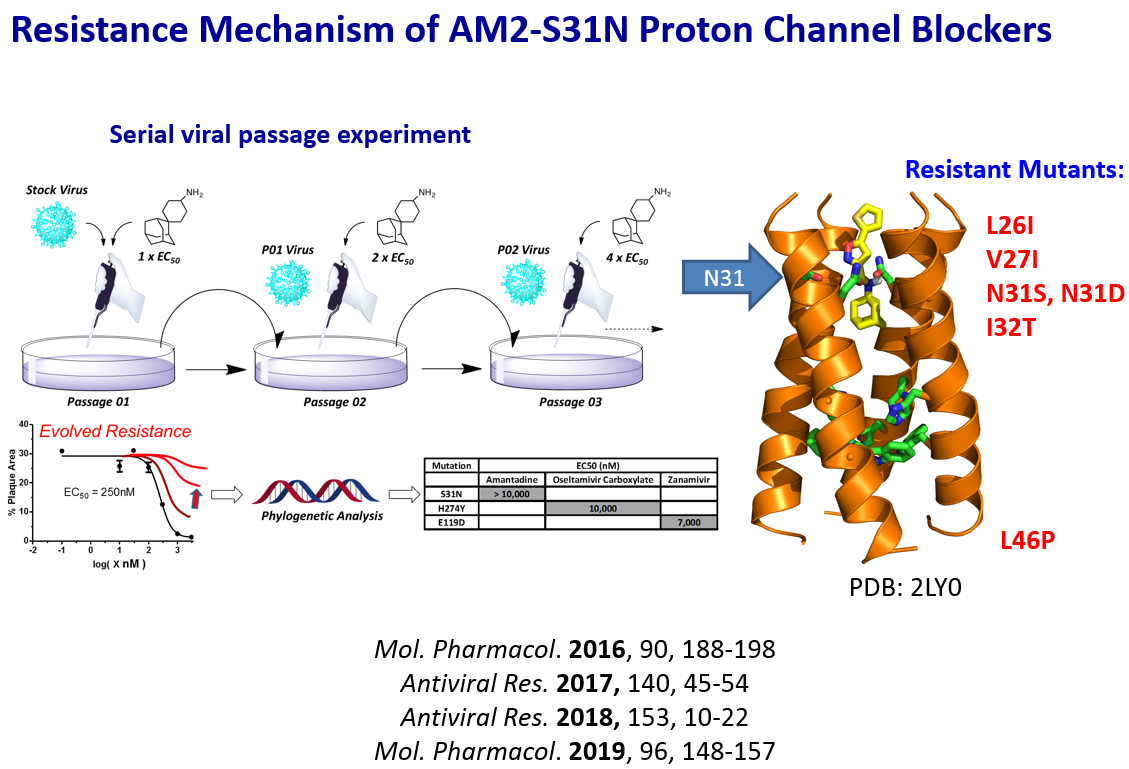
Many anti-infective drugs fail because of drug resistance. Therefore it is essential to elucidate the mechanism of drug resistance and devise corresponding countermeasures before it emerges in clinic. With the antivirals we have developed, we apply them as chemical probes to study the mechanism of drug resistance in serial viral passage experiments, and in turn apply that knowledge to design the next generation of antivirals with a higher genetic barrier to drug resistance. It is astonishing how viruses can evolve and become drug resistance under drug selection pressure. For influenza M2 channel blockers, we found that drug resistance can emerge not only by mutating residues at the drug-binding site, but also residue (L46P) located at allosteric sites that are far away from where the drug binds. To solve the drug resistance issue, we are exploring drug targets that are more conserved as well as combination therapy approach.
Many anti-infective drugs fail because of drug resistance. Therefore it is essential to elucidate the mechanism of drug resistance and devise corresponding countermeasures before it emerges in clinic. With the antivirals we have developed, we apply them as chemical probes to study the mechanism of drug resistance in serial viral passage experiments, and in turn apply that knowledge to design the next generation of antivirals with a higher genetic barrier to drug resistance. It is astonishing how viruses can evolve and become drug resistance under drug selection pressure. For influenza M2 channel blockers, we found that drug resistance can emerge not only by mutating residues at the drug-binding site, but also residue (L46P) located at allosteric sites that are far away from where the drug binds. To solve the drug resistance issue, we are exploring drug targets that are more conserved as well as combination therapy approach.